About iMPI
Non-coding RNAs (ncRNAs) have been recognized not to equip with coding ability for a long time. However, recent advances challenge this widely accepted recognition of ncRNAs, as numerous microprotein-encoding small open reading frames (sORFs) have been discovered in various ncRNAs, such as pre-miRNA, circular RNA, and long non-coding RNA. Introns, another major class of ncRNA, comprise about 40% on average of the total length of genes, and have once been regarded as genome wastes. A growing body of literature has demonstrated that introns would be unspliced and retained in mature mRNA. Then, we inquire whether the retained introns contain sORFs to encode microproteins.
Therefore, in order to systematically explore the translatable introns, we performed genome-wide searching to identify potential encodable intron sORFs (iORFs), and validated the iORF-encoded microproteins using large-scale proteomic mass-spectrometry (MS) data. Finally, we developed the iMPI database to provide an accessible source of iORF-encoded microproteins for researchers.
Genome-wide searching located 209,091 introns in human GRCh38 genome, among which 15,975 were identified to be with high coding potential. Finally, 4,751 introns containing a total of 5,832 iORFs were demonstrated to be potentially translatable according to proteomic MS evidence. The iMPI database provides detailed information on the translatable iORFs, including nucleic sequence, peptide sequence, genomic position, coding potential, and information on relevant introns, transcripts and genes. Incorporating protein MS data from over 27 cancer types, the iMPI database provides cancer-specific MS evidence of iORF-encoded microproteins. Besides, the iMPI database contains detailed annotation regarding the subcellular location, post-translational modifications (83,863 phosphorylation sites, 5,141 ubiquitylation sites, 3,574 methylation sites, 3,363 dimethylation sites, 1,996 trimethylation sites, and 735 acetylation site-specific), second structures, surface accessibility and three-dimensional structure of iORF-encoded microproteins.
209,091 introns from human GRCh38 genome
Incorporating protein MS data from over 27 cancer types.
15,975 introns with high coding potential
4,751 introns containing 5,832 iORFs were potentially translatable.
Various Annotation
Integrating detailed annotations of iORF-encoded microproteins.
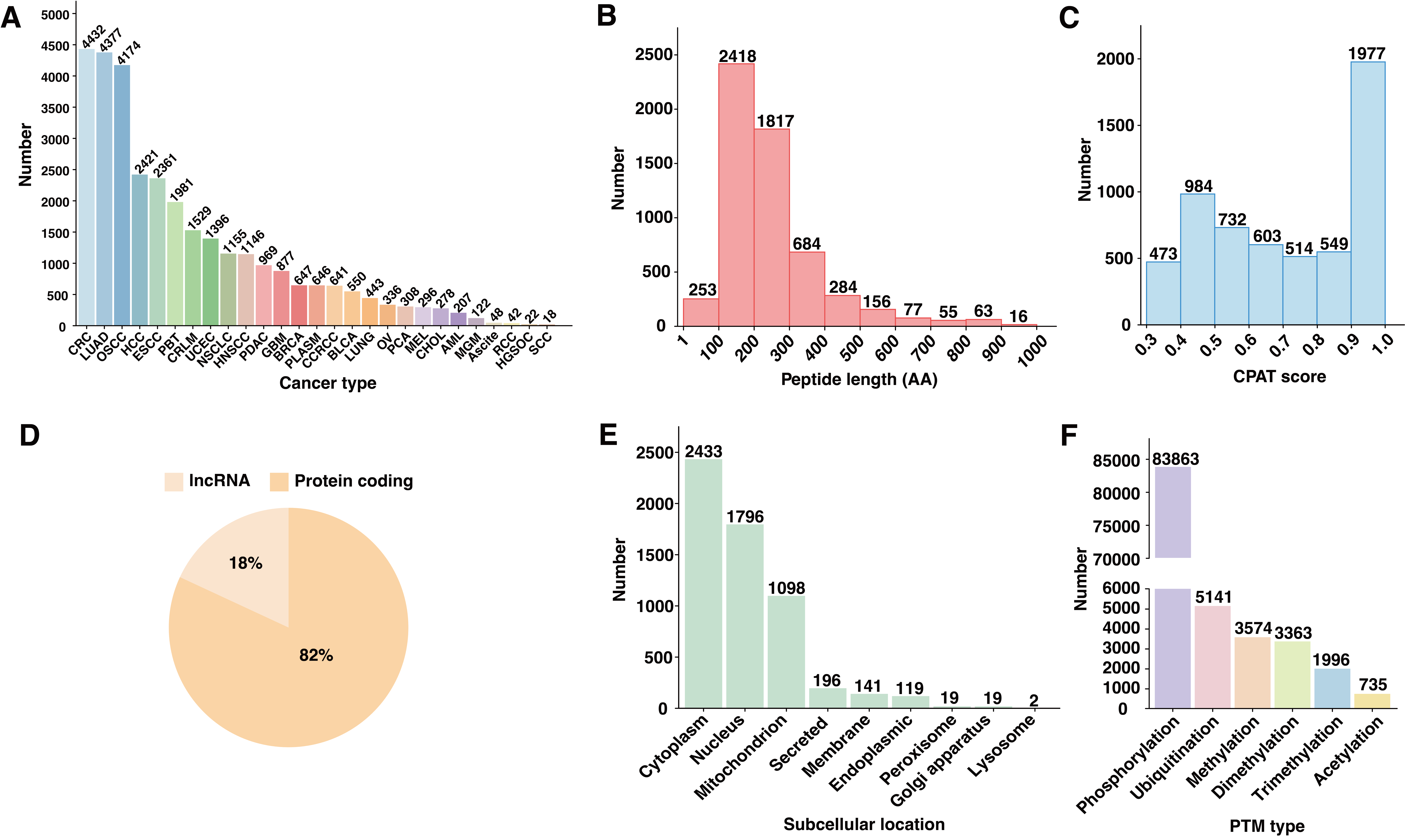
(A) Number of IR-encoded peptides with MS evidence across each cancer type. (B) Distribution spectrum of MPI based on their occurrence across 27 cancer types. MPI were classified into four categories: Tumor-specific (1 cancer type), Shared (2–5 cancer types), Widespread (6–10 cancer types) and Pan-cancer (≥11 cancer types). (C) The inter-cancer overlap of MPI. The vertical axis represents the number of shared MPI among different cancer types, and the horizontal axis lists the number of MPI for each cancer type. The black dots and lines represent the cancer types included in each intersection combination. (D) Distribution of peptide length ranging from 1 to 1,000 amino acids (AA). (E) Distribution of the CPAT score for predicting the coding potential of iORFs. (F) Gene types of introns. (G) Number of IR-encoded peptides localized in different subcellular compartments. (H) Number of PTM sites on IR-encoded peptides.
Mass spectrometry data of specific cancer types used in iMPI
| Cancer type | Abbreviation | Repository | Database identifiers/PMID |
|---|---|---|---|
| Colorectal cancer | CRC | iProx | IPX0000832003 |
| Colorectal cancer | CRC | iProx | IPX0000832010 |
| Hepatocellular carcinoma | HCC | iProx | IPX0000859001 |
| Lung cancer | LUNG | iProx | IPX0001451001 |
| Lung adenocarcinoma | LUAD | iProx | IPX0001804001 |
| Lung cancer | LUNG | Massive | MSV000083969 |
| Bladder cancer | BLCA | Massive | MSV000087370 |
| Melanoma | MEL | PRIDE | PXD001048 |
| Colorectal cancer | CRC | PRIDE | PXD001676 |
| Colorectal cancer | CRC | PRIDE | PXD001794 |
| Non-small cell lung cancer | NSCLC | PRIDE | PXD002612 |
| Ascites | Ascites | PRIDE | PXD003351 |
| Prostate cancer | PCA | PRIDE | PXD003497 |
| Plasma-derived exosomes in Chronic lymphocytic leukemia | PLASMA | PRIDE | PXD004420 |
| Lung cancer | LUNG | PRIDE | PXD004682 |
| Lung cancer | LUNG | PRIDE | PXD004683 |
| Lung cancer | LUNG | PRIDE | PXD004684 |
| Hepatocellular carcinoma | HCC | PRIDE | PXD004873 |
| Breast cancer | BRCA | PRIDE | PXD005214 |
| Hepatocellular carcinoma | HCC | PRIDE | PXD005571 |
| Breast cancer | BRCA | PRIDE | PXD007088 |
| Breast cancer | BRCA | PRIDE | PXD007217 |
| Colorectal liver metastasis | CRLM | PRIDE | PXD008383 |
| Bladder cancer | BLCA | PRIDE | PXD009203 |
| Ascites | Ascites | PRIDE | PXD009382 |
| Colorectal cancer | CRC | PRIDE | PXD009602 |
| Bladder cancer | BLCA | PRIDE | PXD010260 |
| Cholangiocarcinoma | CHOL | PRIDE | PXD010294 |
| Squamous cell carcinoma | SCC | PRIDE | PXD011609 |
| Renal cell carcinoma | RCC | PRIDE | PXD011681 |
| Breast cancer | BRCA | PRIDE | PXD012162 |
| Colorectal cancer | CRC | PRIDE | PXD012254 |
| Breast cancer | BRCA | PRIDE | PXD012431 |
| Meningiomas | MGM | PRIDE | PXD012923 |
| Hepatocellular carcinoma | HCC | PRIDE | PXD013057 |
| Head and neck squamous cell carcinoma; Breast cancer | HNSCC; BRCA | PRIDE | PXD013311 |
| Lung cancer | LUNG | PRIDE | PXD013649 |
| Breast cancer | BRCA | PRIDE | PXD013795 |
| Prostate cancer | PCA | PRIDE | PXD013837 |
| Breast cancer | BRCA | PRIDE | PXD014414 |
| Cholangiocarcinoma | CHOL | PRIDE | PXD017906 |
| Colorectal cancer | CRC | PRIDE | PXD019103 |
| Esophageal squamous cell carcinoma | ESCC | PRIDE | PXD021701 |
| Renal cell carcinoma | RCC | PRIDE | PXD019123 |
| High-Grade Serous Ovarian Cancer | HGSOC | PRIDE | PXD020557 |
| Acute myeloid leukemia | AML | CPTAC | 38232702 |
| Clear cell renal cell carcinoma | CCRCC | CPTAC | 31675502 |
| Glioblastoma | GBM | CPTAC | 33577785 |
| Hepatocellular Carcinoma | HCC | CPTAC | 31585088 |
| Head and neck squamous cell carcinoma | HNSCC | CPTAC | 33417831 |
| Lung adenocarcinoma | LUAD | CPTAC | 32649874 |
| Oral squamous cell carcinoma | OSCC | CPTAC | 28878238 |
| Ovarian serous cystadenocarcinoma | OV | CPTAC | 33086064 |
| Pediatric brain tumors | PBT | CPTAC | 33242424 |
| Pancreatic ductal adenocarcinoma | PDAC | CPTAC | 34534465 |
| Uterine corpus endometrial carcinoma | UCEC | CPTAC | 32059776 |
Annotation sources used in iMPI
| Data type | Resource | Description | URL |
|---|---|---|---|
| Basic information | UniProt | Universal protein resource | https://www.uniprot.org/ |
| Annotation of iORF-encoded microproteins | qPTM | Quantification of post-translational modifications | https://qptm.omicsbio.info/ |
Tools used in iMPI
| Tools | Description | URL |
|---|---|---|
| KMA | An R package that performs intron retention estimation and detection. | https://github.com/pachterlab/kma |
| CPAT | Coding-Potential assessment tool using an alignment-free logistic regression model. | https://github.com/liguowang/cpat |
| MaxQuant | A quantitative proteomics software package designed for analyzing large mass-spectrometric data sets. | https://www.maxquant.org/ |
| AlphaFold | Predicting a protein's 3D structure from its amino acid sequence. | https://alphafold.com/ |
| MULocDeep | A deep-learning framework for protein subcellular and suborganellar localization prediction. | https://github.com/yuexujiang/MULocDeep |
| DSSP | Application to assign secondary structure to proteins. | https://github.com/PDB-REDO/dssp |
MS Data Collection
Collecting MS data from CPTAC Data Portal, iProx, MassIVE, and PRIDE databases.
IR identification
Extracting all introns from the human genome data by KMA (Keep Me Around) algorithm.
Prediction of sequence coding potential
Predicting the coding potential of the introns by CPAT.
Identification of iORF coding ability
Validating the coding potential by performing a database search for protein identification via MaxQuant software.
Annotation of iORF-encoded microproteins
The subcellular location, PTM, tertiary structure, secondary structure, and surface accessibility were predicted and annotated via MULocDeep, qPTM, AlphaFold, and DSSP.
